Precision microbiome insights for families
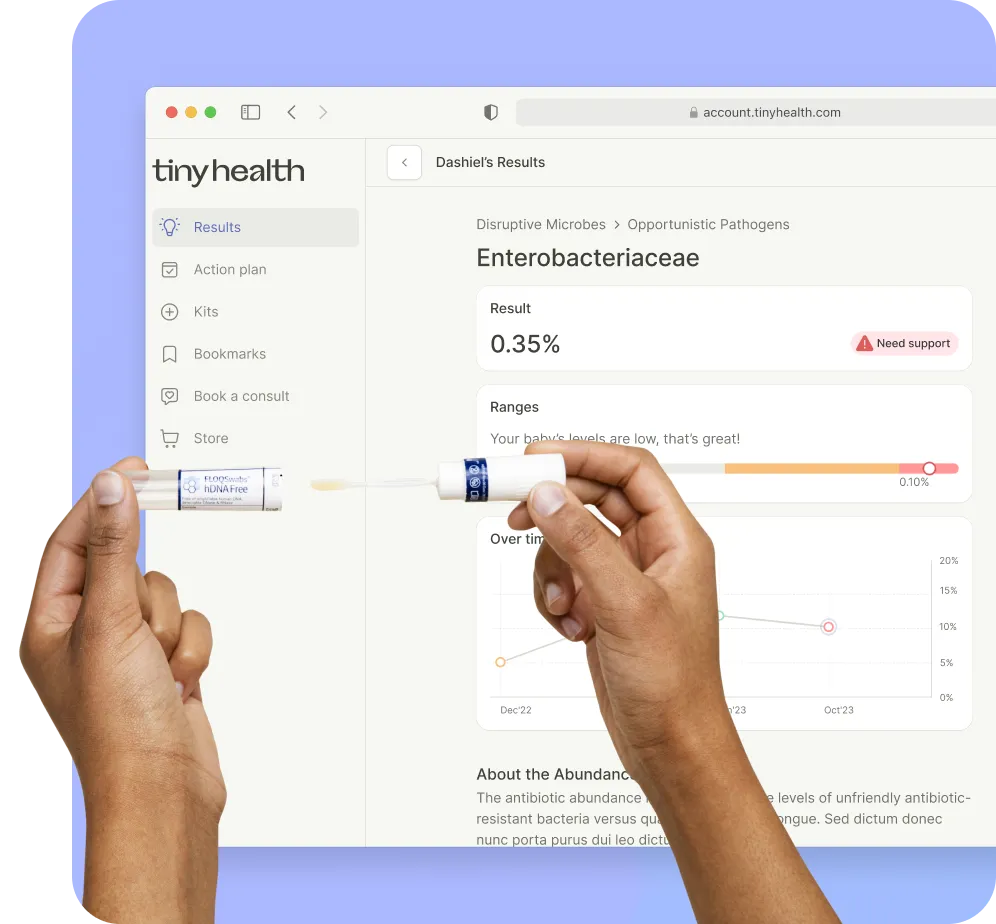
How do we look at the microbiome?

Highest resolution analysis
Tiny Health uses shotgun metagenomics, the gold standard in microbiome research. You get a comprehensive list of all types of microbes found in your gut or vaginal sample, with strain-level precision for key species [1].
Our tests identify over 120,000 bacteria, viruses, fungi, and protozoan parasites—down to a 0.05% relative abundance.
Breakthrough insights
Through functional profiling, we can tell you which microbial genes are present and their abundance levels. This gives us an idea of what the microbes are actually doing.
Our reference ranges and metrics are based on published research and our rapidly growing database of microbiome samples. We have the largest repository of longitudinal pregnancy and baby microbiome samples in the world—helping us advance the science of microbiome health in early life.


Targeted suggestions
Our focus is on prevention, gut health development in the first 1000 days of life, and getting to the root cause of health conditions. We provide actionable advice tailored to your age and unique test results.
No other company delivers our strain-level identification and precision science through at-home tests for the whole family.
Unbiased recommendations
We recommend evidence-based supplements in the market and empower you to pick your own supplements based on the specific strains that may support your specific gut, because we do not believe in a one-size-fits all approach to supplements.

How does our tech compare?
Tiny Health vs. 16s tests
Outdated 16s amplicon (16s) sequencing offers a low resolution view of the microbiome, with a high rate of false positives.

Tiny Health
16s sequencing
Tiny Health vs. qPCR tests
Quantitative polymerase chain reaction (qPCR or PCR) tests are highly specific. But being too specific leads to missing the bigger and more important picture—your overall microbiome health.

Tiny Health
qPCR Tests
Tiny Health vs. mRNA/RNA tests
By analyzing your microbial DNA, we provide you with evidence-based diet, supplement, and lifestyle recommendations. Other tests rely on mRNA (or RNA) which degrades quickly and is easily influenced by factors like time of day and the last thing you ate.

Tiny Health
RNA testing
Tiny Health vs. other metagenomics companies
We’re not the only company to use shotgun metagenomics. But no other company offers deep, strain-level detail paired with Tiny Health’s extensive database. And we aren’t trying to sell you our own branded supplements.
Tiny Health
Metagenomics
Compare microbiome technologies side-by-side
See how our microbiome technology stacks up next to the competition. This chart is best viewed in desktop format.
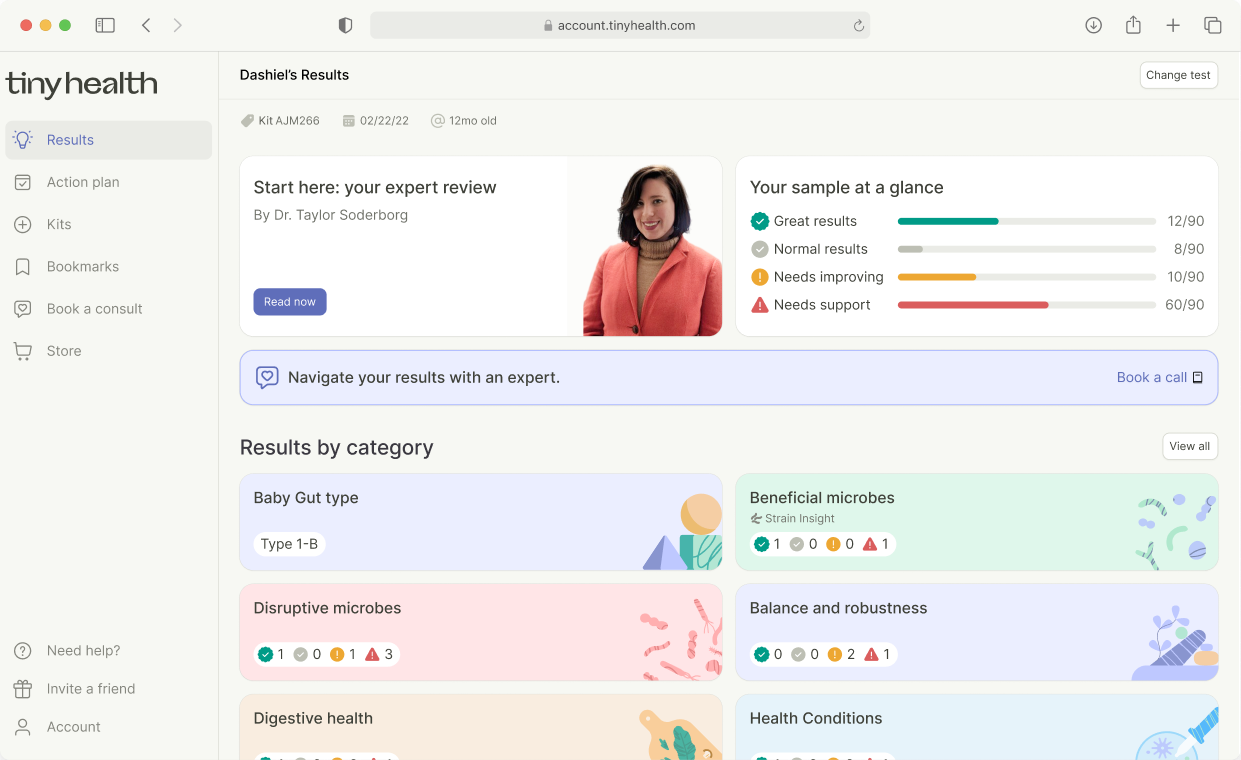
Explore the metrics in our reports
Here are some of the extensive microbiome categories and metrics you’ll find in a Tiny Health report. This list is best viewed in desktop format.
.png)
Feel confident in your next steps toward better health
Tiny Health’s tests bring the most comprehensive microbiome sequencing technology within reach. We support your family’s health journey from before pregnancy through infancy, childhood, and adulthood.
All Tiny Health tests are processed in a CLIA-certified and CAP-certified lab, assuring the highest quality standards.
References
C. Quince, A. W. Walker, J. T. Simpson, N. J. Loman, and N. Segata, “Shotgun metagenomics, from sampling to analysis,” Nat Biotechnol, vol. 35, no. 9, pp. 833–844, Sep. 2017, https://doi.org/10.1038/nbt.3935.
Bashiardes, S., Zilberman-Schapira, G., & Elinav, E. (2016). Use of Metatranscriptomics in Microbiome Research. Bioinformatics and Biology Insights, 10, 19–25. https://doi.org/10.4137/BBI.S34610
Peterson, D., Bonham, K. S., Rowland, S., Pattanayak, C. W., RESONANCE Consortium, & Klepac-Ceraj, V. (2021). Comparative Analysis of 16S rRNA Gene and Metagenome Sequencing in Pediatric Gut Microbiomes. Frontiers in Microbiology, 12, 670336. https://doi.org/10.3389/fmicb.2021.670336
Silverman, J. D., Bloom, R. J., Jiang, S., Durand, H. K., Dallow, E., Mukherjee, S., & David, L. A. (2021). Measuring and mitigating PCR bias in microbiota datasets. PLoS Computational Biology, 17(7), e1009113. https://doi.org/10.1371/journal.pcbi.1009113
Reck, M., Tomasch, J., Deng, Z., Jarek, M., Husemann, P., Wagner-Döbler, I., & COMBACTE Consortium. (2015). Stool metatranscriptomics: A technical guideline for mRNA stabilisation and isolation. BMC Genomics, 16(1), 494. https://doi.org/10.1186/s12864-015-1694-y
Wang, W.-L., Xu, S.-Y., Ren, Z.-G., Tao, L., Jiang, J.-W., & Zheng, S.-S. (2015). Application of metagenomics in the human gut microbiome. World Journal of Gastroenterology: WJG, 21(3), 803–814. https://doi.org/10.3748/wjg.v21.i3.803
Yang, S., & Rothman, R. E. (2004). PCR-based diagnostics for infectious diseases: uses, limitations, and future applications in acute-care settings. The Lancet Infectious Diseases, 4(6), 337–348. https://doi.org/10.1016/S1473-3099(04)01044-8
Yan, Y., Nguyen, L. H., Franzosa, E. A., & Huttenhower, C. (2020). Strain-level epidemiology of microbial communities and the human microbiome. Genome Medicine, 12(1), 71. https://doi.org/10.1186/s13073-020-00765-y
Zhang, X., Li, L., Butcher, J., Stintzi, A., & Figeys, D. (2019). Advancing functional and translational microbiome research using meta-omics approaches. Microbiome, 7(1), 154. https://doi.org/10.1186/s40168-019-0767-6
